Sometimes it’s hard to tell the difference between science and technology ó almost all the time when it has to do with the brain. But this research from MIT that allows for vastly improved scans of the networks inside the brain is too cool to pass up, whether it’s tech, science, or somewhere in between.
Getting up close and personal with neurons and other brain cells is a science that people have been working on for a century and more. Mainly the problem is that they’re so darn small, and packed so tightly, and connect in so many places at once, that it’s hard to tell where anything’s going. We have ways of imaging the brain at various levels, but each is highly limited in its own way.
This new technique addresses several of the main problems. It’s called magnified analysis of proteome, or (conveniently) MAP. The summary from lead researcher Kwanghun Chung makes it sound almost too good to be true.
“We use a chemical process to make the whole brain size-adjustable, while preserving pretty much everything,” Chung says in an MIT news release. “We preserve the proteome (the collection of proteins found in a biological sample), we preserve nanoscopic details, and we also preserve brain-wide connectivity.”

“]Let’s expand that a little. The tissue sample (not living, naturally, and from mice) is suffused with acrylamide polymers. These form a very dense gel – but not so dense that other compounds can’t be sent in, first to attach proteins securely to the gel, then to render them suitable for detection.
At this point, the gel is allowed to expand, increasing to four or five times the original size of the sample. Having something be 800 nanometers across rather than 200 is a big improvement when it comes to imaging and manipulating them, any molecular scientist will tell you. The key feature, though, is that this expansion is somehow non-destructive, preserving the details but blowing them up.
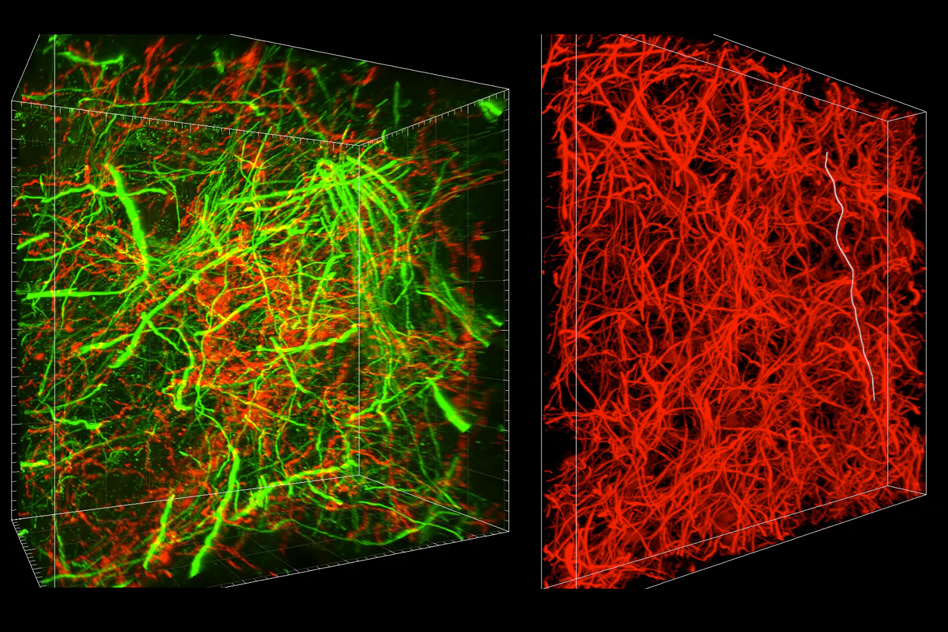
After that, the usual chemical and biological tags can be used to dye and otherwise indicate certain types of cells, reveal where their axons reach to, and so on. A careful 3D scan with a microscope later and you’ve got a fantastically detailed map of quite a large chunk of cortex.
Disrupt 2026: The tech ecosystem, all in one room
Your next round. Your next hire. Your next breakout opportunity. Find it at TechCrunch Disrupt 2026, where 10,000+ founders, investors, and tech leaders gather for three days of 250+ tactical sessions, powerful introductions, and market-defining innovation. Register now to save up to $400.
Save up to $300 or 30% to TechCrunch Founder Summit
1,000+ founders and investors come together at TechCrunch Founder Summit 2026 for a full day focused on growth, execution, and real-world scaling. Learn from founders and investors who have shaped the industry. Connect with peers navigating similar growth stages. Walk away with tactics you can apply immediately
Offer ends March 13.
It’s rather like trying to make out tiny letters written on the surface of a deflated balloon – it’s easier if you just blow it up a bit.
The process is simpler than it sounds, and doesn’t use any exotic materials or radiation, making it potentially very useful for researchers of all stripes and budgets. And it should work on other tissues as well!
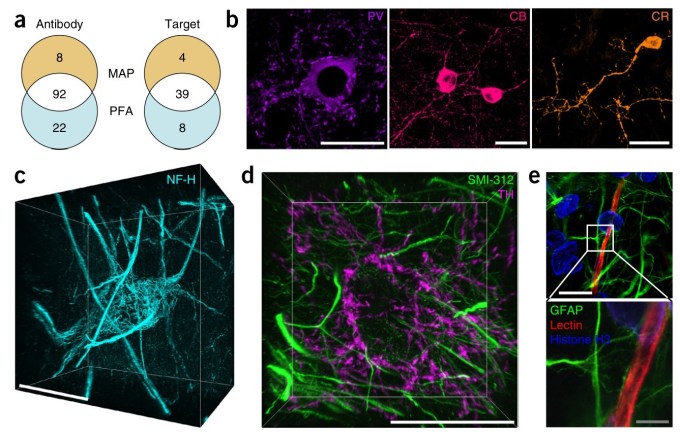
It has the potential to help us better understand real neural networks on both a macro and micro level; models using improved information may help explain how certain aspects of our vision or cognition work, and inform computer models looking to imitate the same.
Lastly, it’s just cool. The seamless melding of biological, chemical, and digital imaging shows how far our tools and knowledge have come in each field. The fact that we can synthesize these disciplines to create something as cohesive as the maps you see above is… well, you read the headline.
Chung et al’s research is published in the journal Nature Biotechnology.